MRF penalty from a phylogeny from a phylo4 object
Usage
# S3 method for class 'phylo4'
mrf_penalty(
object,
model = c("rw1", "ou", "brownian"),
alpha = NULL,
at_tips = NULL,
internal_nodes = TRUE,
delta = FALSE,
...
)Arguments
- object
an R object to create the MRF penalty from.
- model
character; one of
"full"or"individual"indicating if a fully connected graph ("full") or a random effect (random intercepts;"individual") penalty is created.- alpha
numeric; the autoregressive parameter for an OU stochastic process. Should be >1e-5 (alpha = 0 would correspond to a "rw1" model).
- at_tips
character; vector of tip labels to calculate the penalty at. All values in the vector must correpond to tip name labels in the tree.
- internal_nodes
logical; should the internal nodes of the tree be included in the penalty (
TRUE), or just the tips (terminal nodes;FALSE)- delta
numeric or logical; either the numeric value to add to the diagonal of the MRF penalty matrix, or a logical value indicating if such an adjustment should be made. The default is to not alter the diagonal of the penalty matrix.
- ...
arguments passed to other methods.
Examples
#loading the geospiza dataset from phylobase
library(phylobase)
data(geospiza)
#Random-walk (rw1) penalty for both tips and nodes:
pen_rw <- mrf_penalty(geospiza, model = "rw1")
#Same model, but for a reduced number of species
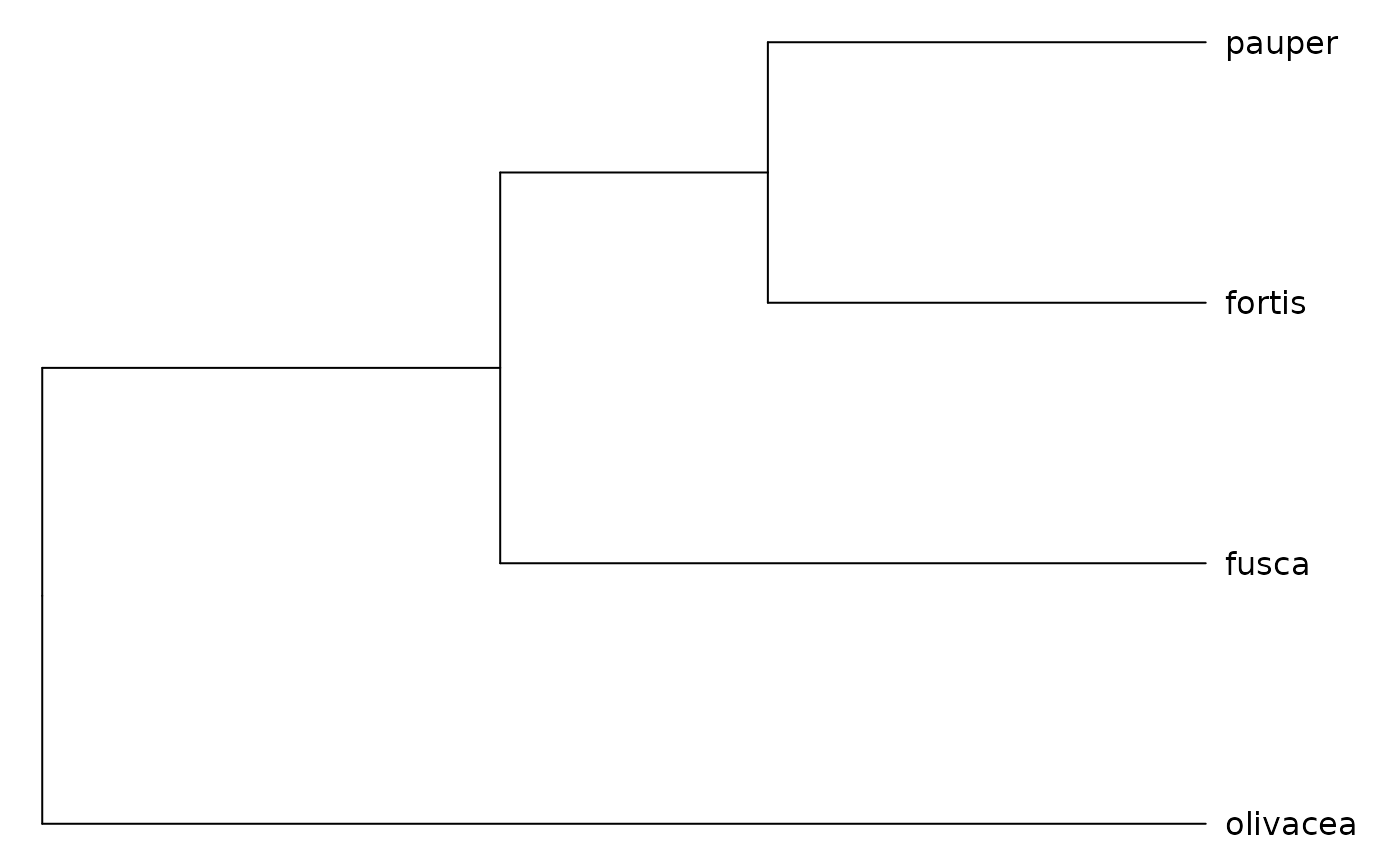
species <- c("fortis", "pauper", "fusca", "olivacea")
pen_rw_subset <- mrf_penalty(geospiza, model = "rw1", at_tips = species)
plot(get_obj(pen_rw_subset))
 #Random-walk penalty matrix for just the tips for all geospiza data:
pen_rw_tips <- mrf_penalty(geospiza, model = "rw1", internal_nodes = FALSE)
pen_rw_tips
#> Markov Random Field penalty
#> Type: tree
#> N : 14
#Ornstein-Uhlenbeck ("ou") process penalty matrix, specifying alpha parameter (autocorrelation)
pen_ou <- mrf_penalty(geospiza, model = "ou", alpha = 1)
#Random-walk penalty matrix for just the tips for all geospiza data:
pen_rw_tips <- mrf_penalty(geospiza, model = "rw1", internal_nodes = FALSE)
pen_rw_tips
#> Markov Random Field penalty
#> Type: tree
#> N : 14
#Ornstein-Uhlenbeck ("ou") process penalty matrix, specifying alpha parameter (autocorrelation)
pen_ou <- mrf_penalty(geospiza, model = "ou", alpha = 1)